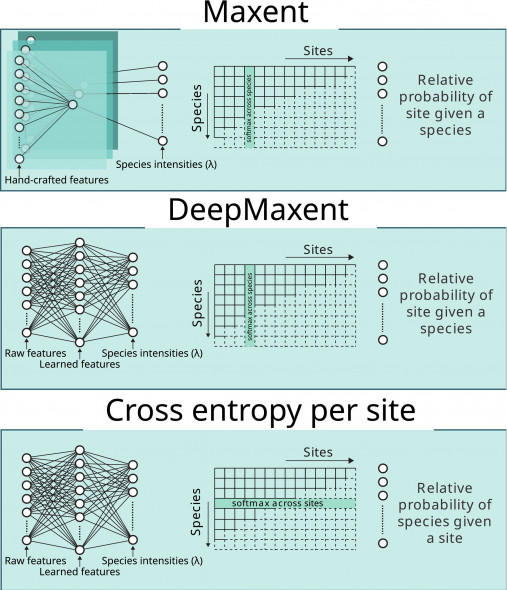
An increase in presence-only (PO) data from citizen science has dramatically enriched biodiversity databases, however their use in species distribution models (SDMs) is still limited by strong sampling biases and the lack of verified absence data. While approaches like Maxent, based on the maximum entropy principle and on Poisson point processes, provide a powerful tool for the creation of SDMs, their dependence on predefined features limits both flexibility and scalability.
In this context, B-Cubed partners analysed how applying the maximum entropy principle to neural networks enhances multi-species distribution models and proposed a method which combines the Maxent principles of maximum entropy with the data-driven feature extraction capabilities of deep learning methods. This new model, DeepMaxent, uses PO data to jointly learn shared latent features and the functions that map from these features to the probability distribution across sites for each species. It provides a flexible foundation to incorporate complex and heterogeneous predictors such as remote sensing imagery, climate projections, ecological context, and multimodal biodiversity data.
The new method relies on a normalised Poisson point-process loss, enabling scalable training with stochastic gradient descent and efficient multi-species modelling while accounting for spatial sampling biases. As a result, DeepMaxent improves predictive accuracy compared to classical Maxent and opens new opportunities for large-scale, multimodal biodiversity monitoring using deep learning.
Read the full article on the topic here.